|
Research interests
Signal transduction mechanisms are instrumental for cells to integrate signals from the external environment and to generate adapted responses. Deregulation of these mechanisms owing to mutations in specific signaling proteins will bring about various pathologies. One of the best and most lethal examples is cancer, which often results from mutations that activate proto-oncogenes (e.g., KRAS, BRAF, PIK3CA) or inactivate tumor suppressor genes (e.g., PTEN, APC, TP53). The objective of the Roux laboratory is to decode the signaling networks within cancer cells to facilitate the development of new anti-cancer treatments.
Using high-throughput proteomic approaches to identify novel effectors of the Ras/MAPK pathway
An important objective of the Roux laboratory is to identify novel effectors of the RAS/MAPK pathway, which is frequently upregulated in human cancer. Melanoma is particularly dependent on the RAS/MAPK pathway, as more than ~ 85% of tumors display mutations affecting one of its components (e.g., BRAF, NRAS, NF1). To better characterize the RAS/MAPK pathway in melanoma and to identify crucial effectors, the Roux laboratory uses several proteomic approaches, including phosphoproteomics and proximity-dependent biotin identification (BioID). Using these methods, they have identified several new effectors of the RAS/MAPK pathway and plan to characterize their biological functions and potential value as novel therapeutic targets.
Identify novel cancer cell vulnerabilities conferred by activated oncogenes Another important objective of the Roux laboratory is to identify novel cancer cell vulnerabilities conferred by activated oncogenes. To better understand the role of oncogenic signaling in tumorigenesis and drug resistance, they have developed several cellular models harboring some of the main activated oncogenes found in human cancers. Using a proteogenomics approach relying on transcriptomics, proteomics and CRISPR-based functional screening, the Roux laboratory seeks to identify potential vulnerabilities associated with the KRAS, PIK3CA, MYC and HER2 oncogenes.
Identify new immunotherapeutic targest in cancer The Roux laboratory seeks to identify new diagnostic tools and immunotherapeutic targets in diverse types of cancer. The use of antibody therapies in human diseases, in particular cancer, is gaining importance, but a major technical challenge is finding the few cell-surface proteins that provide distinguishing marks for specific cancer types. Recent advances in chemoproteomics have begun to overcome these limitations by combining chemical tagging of cell-surface proteins with affinity purification-mass spectrometry (AP-MS). The Roux laboratory has adapted these approaches to simultaneously identify and quantify cell-surface antigens from primary specimens and cancer cell lines. A major objective of the Roux team is to characterize these potential targets and develop antibody-based immunotherapy.
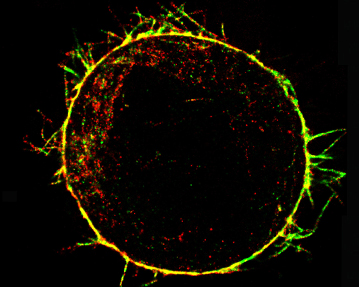
Interplay between mechanical and biological mechanisms in cell cortex assembly The cell cortex is a network of actin, myosin and associated proteins that underlies the plasma membrane and determines cell shape. As such, the cortex plays a role in normal physiology during events involving cell deformation such as mitosis, cytokinesis, and cell locomotion, and in the pathophysiology of diseases such as cancer. Despite its importance, little is known about how the cortex assembles, how it is tethered to the cell membrane, and how its mechanical properties are regulated. The aim of this project is to uncover the molecules involved in cortex assembly, understand their regulation, determine how cortex mechanical properties relate to cortical ultrastructure and proteic composition, and explore how cortical mechanics in turn influence molecular concentrations and activities. This interdisciplinary project, in collaboration with Guillaume Charras (University College London), Ewa Paluch (University of Cambridge), and Guillaume Romet-Lemonne (Institut Jacques Monod), will enable us to rationally link proteic composition, ultrastructure, mechanics and regulation of the cortex to one another.
|
Normal human mammary epithelial cells (actin filaments are stained in red, DNA in blue)
Breast cancer cells with hyperactivated MAPK signalling (as above, with the Gab2 oncogene stained in green)
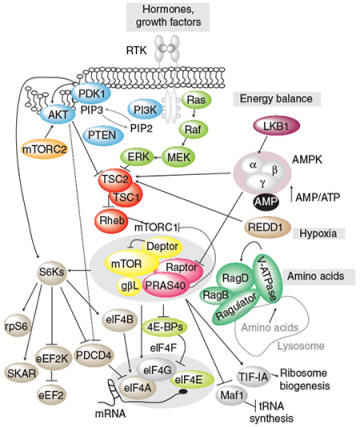
Schematic of the signal transduction mechanisms stimulated by hormones and growth factors (from Roux and Topisirovic, CSHPB 2012)
Actin cytoskeleton of a rounded cell in mitosis viewed using an electron microscope (From Serres et al., Dev Cell 2020) |
.jpg)
.jpg)